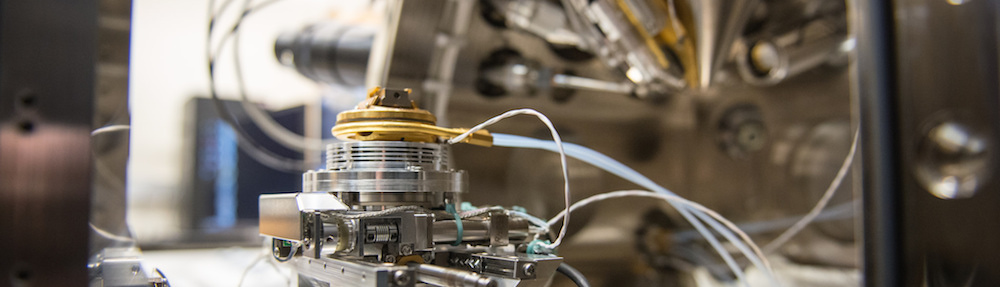
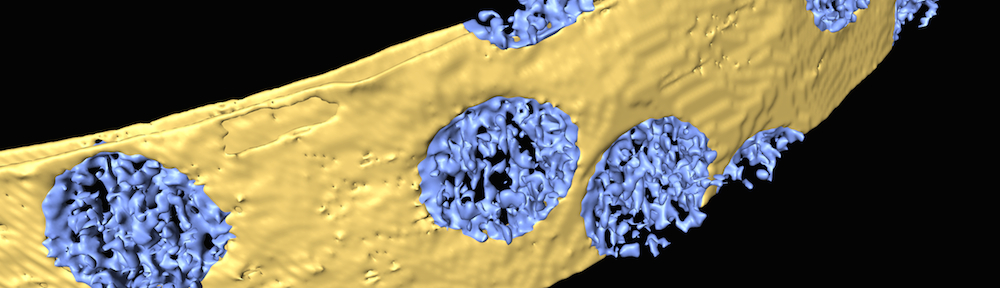
The Villa Lab develops tools to observe macromolecular complexes in their natural environment, the cell. Our ultimate goal is to unveil the structural dynamics of these complexes while they go about their everyday life. We combine cell biology and electron microscopy to generate data, and we use image processing and physical modeling to understand these data.
EPI
Being part of a dream team of basic and translational scientists working toward a common goal is exhilarating. Thanks HHMI for support! #phagetherapy #phagenucleus #AntibioticResistance
Digvijay’s cell paper
Digvijay’s work on the in situ yeast NPC has now been published after peer review #openaccess. This work was a massive collaboration with the labs of Chris Akey, Mike Rout, Andrej Sali, and Steve Ludke. The article describes comprehensive NPC structures in vitro and in situ , and shows the existence of three NPC forms co-exist in the same cell.
welcome Tamar
Tamar Basiashvili is joining our lab as a graduate student. Welcome Tamar!
Matt’s DC paper
Matt’s paper showing how deconvolution can help cryo-ET data is now out after peer review #openaccess. This paper is a product of a wonderful long-standing collaboration with John Sedat.
HHMI
Elizabeth is selected as a Howard Hughes Medical Institute (HHMI) Investigator. This wonderful opportunity which was only possible through the work of lab members through the years, will allow us to continue to pursue high-risk research. We’re so excited!